Research:
BREAST CANCER METASTASES
Additional research on metastatic cancer is necessary to decrease the number of patients dying from breast cancer. We have focused on identifying druggable mediators of metastases to the brain and bone. Our sequencing analysis of intra-patient paired primary and metastatic lesions to the brain lead to a number of clinically relevant findings, including ERBB2/HER2 alterations in approximately 20% of ERBB2/HER2-negative cases (Priedigkeit et al., 2017a). In collaboration with Dr Leonie Young’s team (RCS, Dublin), we identified recurrent gain of RET expression in many brain metastases (Vareslija et al., 2019), and we are currently dissecting the underlying mechanisms and potential clinical opportunities.
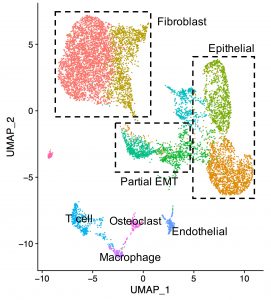
Bone metastases occur in the majority of patients with ER+ breast cancer. Using capture RNA sequencing approaches, we showed that decade-old and decalcified specimens can be utilized for analysis of gene expression. We identified a number of longitudinal expression alterations of clinically actionable genes, particularly in the CDK/Rb/E2F and FGFR signaling pathways (Priedigkeit et al., 2017b). In collaboration with Drs Weiss and Watters (UPMC HCC) we are expanding these studies applying single cell sequencing approaches in order to understand intratumor heterogeneity.
Given our interest in lobular diseases, we aim to identify drivers of ILC metastases to unique sites, such as the ovary, and the gastrointestinal tract. For these studies, we are using clinical samples, mouse models and in vitro approaches. We recently finished a characterization of orbital and periorbital metastases in patients with ILC, identifying frequent contralateral orbital and co-occurring CNS metastases (Blohmer et al, in revision). We have also identified FGFR4 overexpression and (rare) mutations enriched in ILC metastases (Levine et al., 2019), a finding that we are following up with further mechanistic analyses, and studies to assess potential therapeutic targeting.
Finally, we are leading a large multi-disciplinary team using clinical material from breast cancer patients who donate their bodies to research, frequently called “Rapid Autopsy”, a form of organ donation following death. These generous donations will help us to decipher mechanisms of breast cancer organ tropism, evolution and drivers of metastases. Through analysis of samples from the Rapid Autopsy program we have previously identified an ER fusion (Hartmaier et al., 2018), described in more detail in “Endocrine Resistance” section above.
Endocrine Resistance | Invasive Lobular Breast Cancer | Precision Medicine
Hartmaier, R. J., Trabucco, S. E., Priedigkeit, N., Chung, J. H., Parachoniak, C. A., Vanden Borre, P., Morley, S., Rosenzweig, M., Gay, L. M., Goldberg, M. E., et al. (2018). Recurrent hyperactive ESR1 fusion proteins in endocrine therapy-resistant breast cancer. Ann Oncol 29, 872-880.
Levine, K. M., Priedigkeit, N., Basudan, A., Tasdemir, N., Sikora, M. J., Sokol, E. S., Hartmaier, R. J., Ding, K., Ahmad, N. Z., Watters, R. J., et al. (2019). FGFR4 overexpression and hotspot mutations in metastatic ER+ breast cancer are enriched in the lobular subtype. NPJ Breast Cancer 5, 19.
Priedigkeit, N., Hartmaier, R. J., Chen, Y., Vareslija, D., Basudan, A., Watters, R. J., Thomas, R., Leone, J. P., Lucas, P. C., Bhargava, R., et al. (2017a). Intrinsic Subtype Switching and Acquired ERBB2/HER2 Amplifications and Mutations in Breast Cancer Brain Metastases. JAMA Oncol 3, 666-671.
Priedigkeit, N., Watters, R. J., Lucas, P. C., Basudan, A., Bhargava, R., Horne, W., Kolls, J. K., Fang, Z., Rosenzweig, M. Q., Brufsky, A. M., et al. (2017b). Exome-capture RNA sequencing of decade-old breast cancers and matched decalcified bone metastases. JCI Insight 2.
Vareslija, D., Priedigkeit, N., Fagan, A., Purcell, S., Cosgrove, N., O’Halloran, P. J., Ward, E., Cocchiglia, S., Hartmaier, R., Castro, C. A., et al. (2019). Transcriptome Characterization of Matched Primary Breast and Brain Metastatic Tumors to Detect Novel Actionable Targets. J Natl Cancer Inst 111, 388-398.

